In Situ Hybridization in Cytogenetics
Introduction
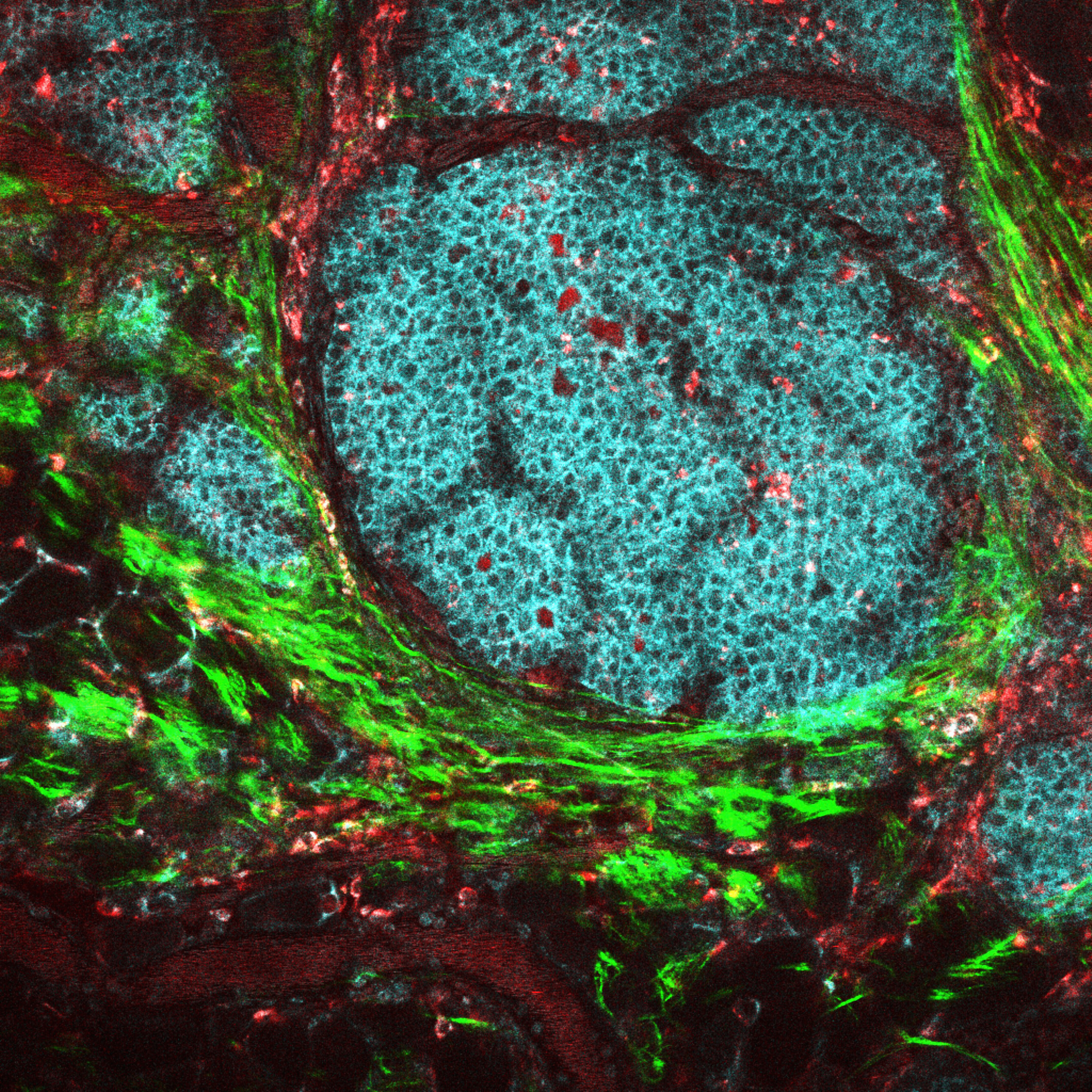
In situ hybridization (ISH) is a powerful technique used in cytogenetics to detect and localize specific DNA sequences directly on chromosomes. By allowing precise mapping of genetic material, ISH has become an essential tool in modern molecular biology, medical diagnostics, and genomic research.
First developed in the late 1960s, this method has evolved significantly and now enables the identification of unique DNA sequences within complex genomes.
Principle of In Situ Hybridization
ISH is based on the hybridization of a labeled DNA probe with its complementary sequence on chromosomal DNA.
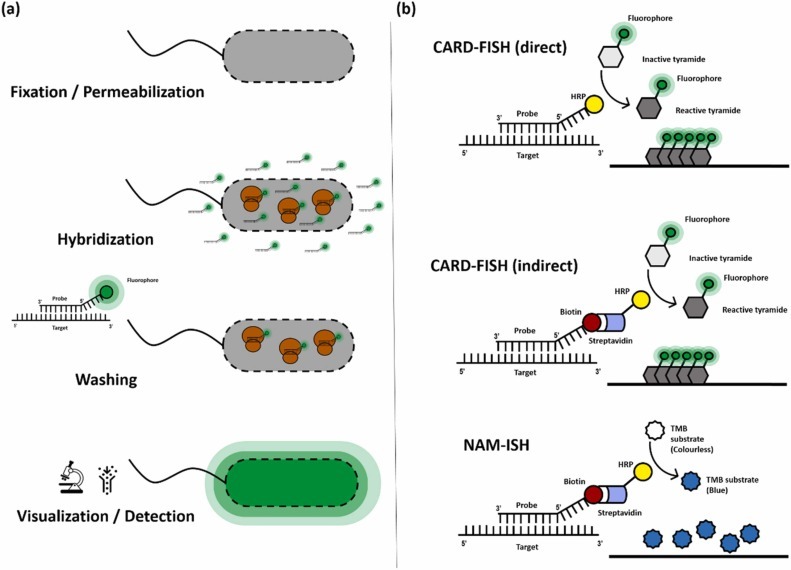
The process involves three main steps:
- Preparation of a labeled DNA probe
- Hybridization with chromosomal DNA
- Detection and analysis of hybridization signals
This approach allows researchers to visualize the exact chromosomal location of a gene or DNA fragment.
Preparation of DNA Probes
DNA probes are typically fragments of DNA cloned into vectors and labeled for detection. Traditionally, probes were labeled using radioactive isotopes such as tritium.
Key steps include:
- Labeling DNA using techniques like nick translation
- Denaturation of the probe at high temperature (~70°C)
- Preparation in a hybridization solution containing agents like formamide and dextran sulfate
These components enhance probe binding efficiency and specificity.
Hybridization Process
Chromosomal preparations are obtained from cells in metaphase, where chromosomes are most visible.
Before hybridization:
- Chromosomal DNA is denatured to make it single-stranded
- RNA is removed to prevent non-specific binding
- Slides are treated to optimize accessibility of DNA
The probe is then applied and incubated (typically at ~42°C) to allow binding with complementary sequences.
Detection and Autoradiography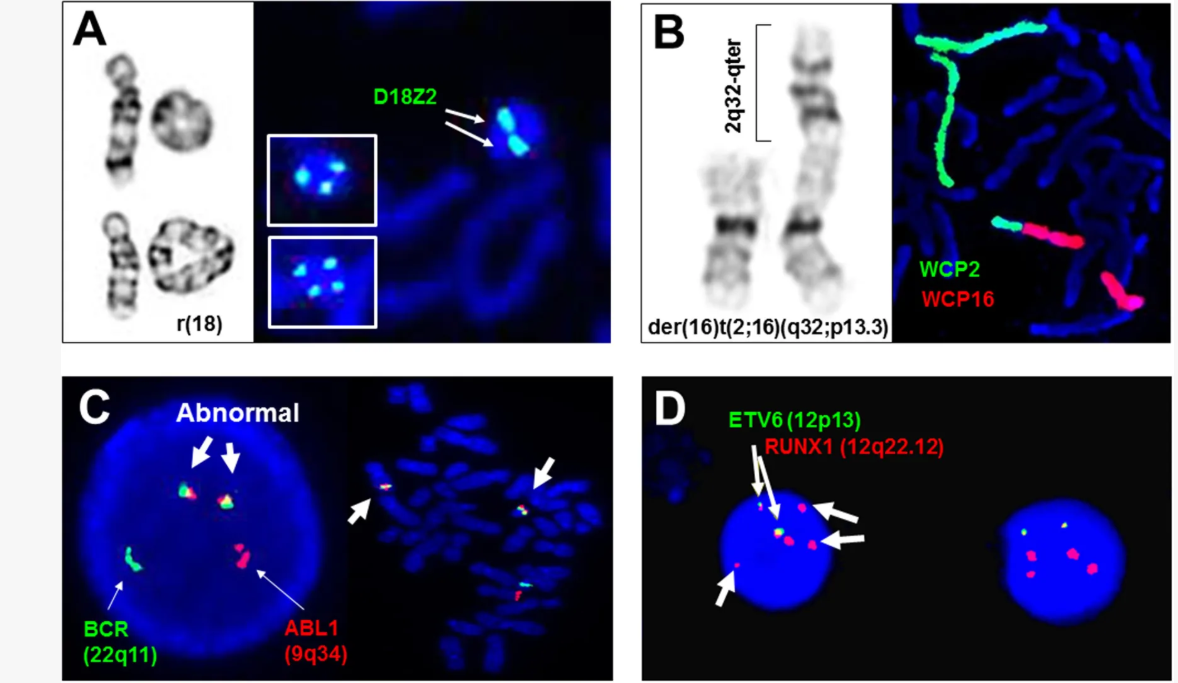
After hybridization, detection is performed using autoradiography when radioactive probes are used.
This involves:
- Covering slides with a photographic emulsion
- Exposure for several days to detect radioactive signals
- Visualization of silver grains indicating hybridization sites
Each signal corresponds to a specific chromosomal region where the probe has bound.
Interpretation of Results
Microscopic analysis allows identification of labeled chromosomal regions.
Key steps include:
- Observing well-spread metaphase chromosomes
- Mapping signals onto chromosomal banding patterns
- Constructing karyotype diagrams
Clusters of signals indicate strong hybridization and precise gene localization.
Gene Mapping Using ISH
ISH has revolutionized gene mapping by providing a rapid and direct method for locating genes on chromosomes.
Unlike traditional methods such as:
ISH allows:
- High-resolution localization
- Direct visualization of gene positions
- Improved accuracy in genomic mapping
For example, important genes on the human X chromosome have been precisely mapped using ISH, improving our understanding of genetic disorders.
Applications in Evolutionary Biology
ISH is also widely used in comparative genomics.
Because some genes are conserved across species, the same probe can hybridize with DNA from different organisms, such as humans and mice. This allows:
- Identification of homologous chromosomal regions
- Study of chromosomal rearrangements
- Understanding of genome evolution
Such analyses are essential for developing animal models of human diseases.
Role in Cytogenetic Diagnosis
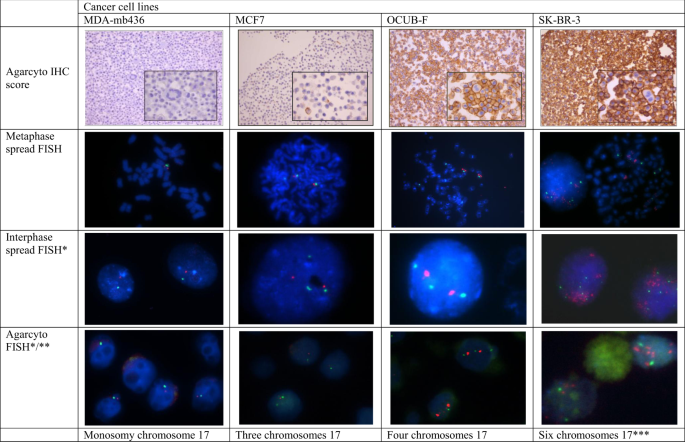
ISH plays a critical role in diagnosing chromosomal abnormalities, including:
- Trisomies and tetrasomies
- Small supernumerary chromosomes
- Structural rearrangements (translocations, deletions)
It is particularly useful when traditional cytogenetic techniques cannot clearly identify chromosomal structures.
ISH also enables detection of submicroscopic abnormalities, including small DNA segments that are not visible under a microscope.
Applications in Cancer Research
ISH has become a key tool in cancer genetics.
It helps detect:
- Activation of oncogenes
- Gene amplification
- Chromosomal rearrangements associated with tumors
For example, specific chromosomal translocations involving oncogenes are linked to certain cancers, such as Burkitt lymphoma.
Additionally, ISH can identify gene amplification patterns, which appear as:
- Double minute chromosomes (DMs)
- Homogeneously staining regions (HSRs)
These features are associated with increased oncogene expression.
Advances in Non-Radioactive Probes
Modern ISH techniques increasingly use non-radioactive probes labeled with:
- Fluorescent dyes
- Enzymatic markers
These methods offer several advantages:
- Faster detection (within 24 hours)
- Improved safety
- Better reproducibility
However, their resolution may still be lower compared to traditional radioactive probes for very small DNA sequences.
Conclusion
In situ hybridization is a cornerstone technique in cytogenetics and molecular biology. It enables precise localization of DNA sequences, supports gene mapping, and plays a vital role in diagnosing genetic disorders and studying cancer.
By bridging the gap between cytogenetics and molecular biology, ISH continues to advance our understanding of genome structure, function, and evolution.
Recent Posts
-
Randomly Amplified Polymorphic DNA (RAPD)
Introduction The analysis of genetic variation at the DNA level has become a fundamental approach in …1st Apr 2026 -
Role of Cytogenetics and Molecular Cytogenetics in Diagnosing Genetic Imbalances
Introduction Over the past five decades, cytogenetics and molecular cytogenetics have transformed t …1st Apr 2026 -
In Situ Hybridization in Cytogenetics
Introduction In situ hybridization (ISH) is a powerful technique used in cytogenetics to detect and …31st Mar 2026