Randomly Amplified Polymorphic DNA (RAPD)
Introduction
The analysis of genetic variation at the DNA level has become a fundamental approach in modern molecular biology. Several techniques such as RAPD (Randomly Amplified Polymorphic DNA), AFLP, RFLP, SSR, and microsatellites are widely used to study genetic diversity. Among these, RAPD has gained significant attention since the 1990s due to its simplicity, speed, and cost-effectiveness.
RAPD is a PCR-based molecular marker technique that enables the detection of genetic polymorphisms without prior knowledge of DNA sequences. By using short, arbitrary primers, RAPD amplifies multiple DNA fragments across the genome, generating unique banding patterns that reflect genetic variation.
Principle of RAPD
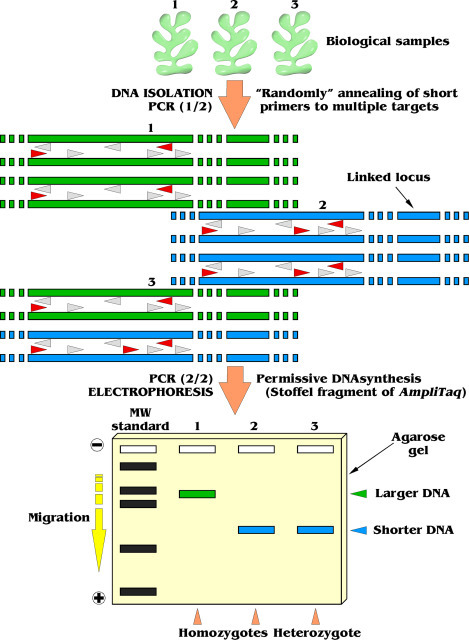
RAPD relies on the amplification of random DNA segments using a single short oligonucleotide primer (typically 8–12 nucleotides) under low annealing temperatures.
During PCR:
- The primer binds at multiple random sites in the genome
- DNA fragments are amplified only when two primer-binding sites are within amplifiable distance
- The resulting fragments are separated using agarose gel electrophoresis, producing a DNA fingerprint
The banding pattern depends on sequence complementarity between the primer and template DNA, allowing detection of polymorphisms across multiple loci simultaneously.
Methodology of RAPD
1. DNA Extraction
Genomic DNA is extracted from biological samples (e.g., blood, tissue) using standard protocols such as:
- Proteinase K digestion
- Phenol–chloroform purification
2. DNA Quality and Quantification
DNA integrity and purity are assessed using:
- UV spectrophotometry (OD260/OD280 ratio)
- Agarose gel electrophoresis
3. Primer Selection
- Short arbitrary primers (usually 10 bp, GC-rich) are used
- No prior genomic sequence information is required
- Multiple primers can be tested to maximize polymorphism detection
4. PCR Amplification
RAPD PCR involves three main steps:
- Denaturation (≈94°C): DNA strands separate
- Annealing (≈35–42°C): Primer binds to random sites
- Extension (≈72°C): DNA polymerase synthesizes new strands
The reaction mixture typically includes:
- DNA template
- Primers
- dNTPs
- MgCl₂
- Taq DNA polymerase
- Reaction buffer
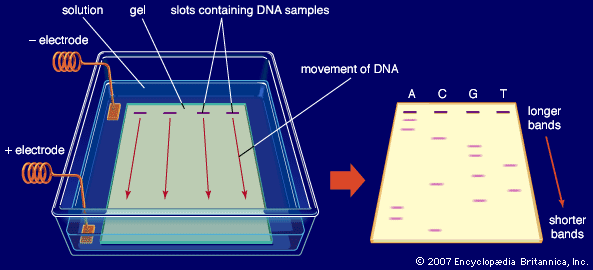
5. Gel Electrophoresis
Amplified DNA fragments are separated on agarose gel and visualized under UV light after staining (e.g., ethidium bromide). The resulting banding pattern represents the genetic profile.
Data Analysis in RAPD Studies
RAPD data are analyzed using statistical and computational tools to evaluate genetic variation:
- Genetic diversity indices: allele frequency, Shannon’s index
- Neutrality tests: Ewens–Watterson test
- Population differentiation: AMOVA (Analysis of Molecular Variance)
- Gene flow estimation: Nm values
- Cluster analysis: UPGMA dendrograms
These analyses help quantify genetic relationships within and between populations.
Advantages of RAPD
RAPD remains popular due to several key benefits:
- No requirement for prior DNA sequence information
- Rapid and cost-effective technique
- Requires only small amounts of DNA
- Simultaneous analysis of multiple loci
- Suitable for poorly characterized genomes
- High potential for detecting polymorphism
Additionally, RAPD markers can be cloned and further analyzed for advanced genetic studies.
Limitations of RAPD
Despite its advantages, RAPD has several technical constraints:
- Dominant markers: cannot distinguish homozygous from heterozygous genotypes
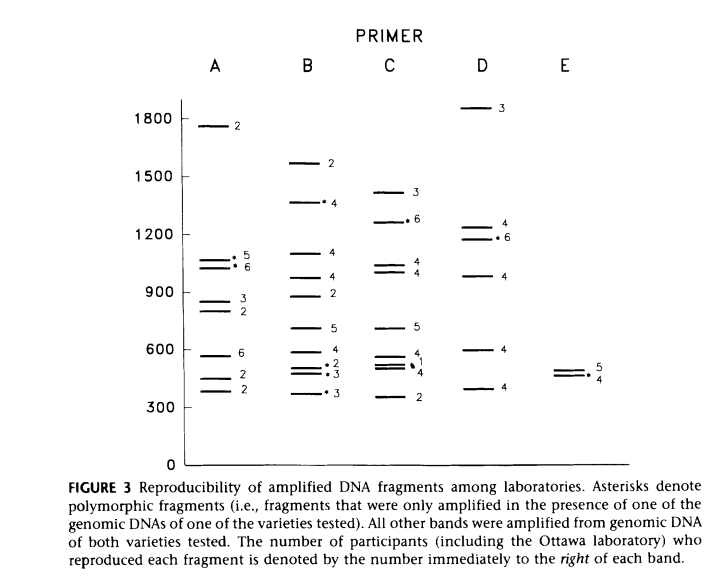
- Reproducibility issues: sensitive to PCR conditions and DNA quality
- Band ambiguity: co-migrating fragments may not represent identical sequences
- Limited specificity: unknown identity of amplified fragments
Careful optimization of experimental conditions is essential to ensure reliable results.
Applications of RAPD
1. Genetic Mapping
RAPD markers can be used to identify polymorphic DNA regions and assist in gene mapping, complementing traditional methods like RFLP.
2. Marker-Assisted Selection
RAPD enables identification of markers linked to specific traits, supporting breeding programs without full genome mapping.
3. Genetic Diversity Analysis
Widely used to evaluate:
- Intraspecific variation
- Population structure
- Phylogenetic relationships
4. Population and Evolutionary Genetics

RAPD helps in studying:
- Genetic relationships among populations
- Evolutionary patterns
- Conservation genetics
5. Plant and Animal Breeding
The technique supports:
- Detection of inbreeding levels
- Identification of elite genotypes
- Improvement of quantitative traits
6. Microbial and Species Identification
RAPD is effective for:
- Strain differentiation
- Detection of genetic heterogeneity
- Identification of species-specific markers
Reproducibility Considerations
Although RAPD is simple and fast, reproducibility depends on strict standardization of:
- DNA quality and concentration
- Primer selection
- PCR conditions
Variations in these parameters can significantly affect banding patterns, making protocol consistency essential.
Conclusion
RAPD is a powerful, rapid, and cost-efficient molecular marker technique widely used in genetic research. It plays a crucial role in genetic diversity analysis, breeding programs, and population genetics, particularly when genomic information is limited.
Despite limitations related to reproducibility and marker dominance, RAPD remains highly valuable, especially in resource-limited laboratories. Its ability to generate multiple markers quickly makes it an important tool in modern molecular genetics and biodiversity studies.
Recent Posts
-
Randomly Amplified Polymorphic DNA (RAPD)
Introduction The analysis of genetic variation at the DNA level has become a fundamental approach in …1st Apr 2026 -
Role of Cytogenetics and Molecular Cytogenetics in Diagnosing Genetic Imbalances
Introduction Over the past five decades, cytogenetics and molecular cytogenetics have transformed t …1st Apr 2026 -
In Situ Hybridization in Cytogenetics
Introduction In situ hybridization (ISH) is a powerful technique used in cytogenetics to detect and …31st Mar 2026


